Research
Our lab works at the interface of engineering and biology to engineer microbes and consortia with novel functions. We are especially interested in deciphering how “unwieldy” microbes in the environment perform extraordinary tasks - many of these microbes have no available genomic sequence and are exceptionally difficult to manipulate.
Genetic and Cellular Engineering of Anaerobic Gut Fungi
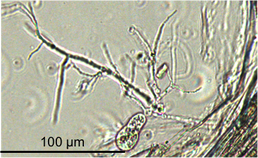
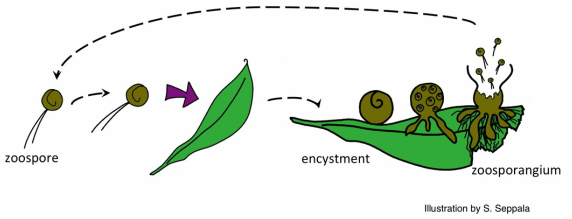
Anaerobic gut fungi colonizing wheat straw.
Chemicals derived from lignocellulose (plants) are an attractive alternative to petroleum-sourced chemicals. The most tractable approach to harness the energy contained within biomass is to break down the cellulosic polymer into simple sugars that can be converted via microbial fermentation into energy-rich organic fuels such as ethanol or butanol. However, the complex structure of cellulose requires the synergistic action of several different enzymes for complete hydrolysis; these enzymes are limited to those made by model cellulolytic microbes, which are ubiquitous and easy to culture, yet enzyme catalytic abilities are limited and their production/purification is costly.
Towards sustainable chemical production from biomass, much may be learned by studying nature – particularly environments where routine, high-efficiency lignocellulose breakdown regularly occurs. The microbial world holds a wealth of lignocellulolytic microbes (and enzymes) left undiscovered by conventional means; for example, the digestive tract of large herbivores has long been recognized as the best locale for cellulosic biomass breakdown. Anaerobic fungi that reside within the digestive tract of large herbivores are among the most efficient and robust digesters of lignocellulosic material known in nature owing to their invasive, filamentous growth habit. Despite the powerful cellulose-degrading capacity of these fungi, remarkably little is known about the spectrum of cellulolytic enzymes produced by these organisms. We are developing new tools to identify novel cellulolytic enzymes that originate from anaerobic fungi, and to genetically engineer them for lignocellulosic bioprocessing. In recent years, our group has developed unique skills to isolate, characterize, and cultivate these organisms that combines classic anaerobic microbiology with RNA-Seq approaches. This information will guide in genetic transformation and reprogramming strategies to be applied to the fungi, which will allow their metabolic pathways to be tuned according to need.
Towards sustainable chemical production from biomass, much may be learned by studying nature – particularly environments where routine, high-efficiency lignocellulose breakdown regularly occurs. The microbial world holds a wealth of lignocellulolytic microbes (and enzymes) left undiscovered by conventional means; for example, the digestive tract of large herbivores has long been recognized as the best locale for cellulosic biomass breakdown. Anaerobic fungi that reside within the digestive tract of large herbivores are among the most efficient and robust digesters of lignocellulosic material known in nature owing to their invasive, filamentous growth habit. Despite the powerful cellulose-degrading capacity of these fungi, remarkably little is known about the spectrum of cellulolytic enzymes produced by these organisms. We are developing new tools to identify novel cellulolytic enzymes that originate from anaerobic fungi, and to genetically engineer them for lignocellulosic bioprocessing. In recent years, our group has developed unique skills to isolate, characterize, and cultivate these organisms that combines classic anaerobic microbiology with RNA-Seq approaches. This information will guide in genetic transformation and reprogramming strategies to be applied to the fungi, which will allow their metabolic pathways to be tuned according to need.
Synthetic Anaerobic Consortia for Bioproduction and Model Development
Anaerobic microbes have evolved to work together in complex communities that decompose and recycle carbon biomass throughout the Earth – from our guts to landfills and compost piles. Compared to microbes that thrive in the presence of oxygen, anaerobes are woefully understudied; they represent a vast, untapped resource for novel enzymes that degrade woody biomass into sugars, as well as natural products that could find use as new drugs. Despite their importance, little information exists to parse the role of each microbial member within their dynamic community. In particular, their metabolic “connection points” are vague, their secreted products are uncharacterized, and the enzyme machinery that they use to extract sugars from crude biomass is elusive. By understanding these natural systems, engineers can build new technologies that extract sugar from renewable plant biomass for conversion into commodity chemicals – an exciting alternative to producing chemicals from petroleum feedstocks. Similarly, an understanding of how anaerobic microbes “defend themselves” from neighbors could translate into new antimicrobial compounds for therapeutic use.
To address these knowledge gaps, we have pioneered new techniques to isolate anaerobes from biomass-rich environments (e.g. guts and fecal materials of herbivores) and build synthetic consortia to uncover their interdependencies. We are developing new tools to construct synthetic microbial communities based on how natural anaerobic consortia interact. Our ultimate goal is to develop systems-level tools to evaluate and direct anaerobic microbial interaction, such that carbon flux can be engineered within these communities by compartmentalizing breakdown and production across microbes. Forthcoming applications of this research include engineering anaerobic digestors, anaerobic production of fuels and commodity chemicals from biomass, and identification of natural products from gut anaerobes.
To address these knowledge gaps, we have pioneered new techniques to isolate anaerobes from biomass-rich environments (e.g. guts and fecal materials of herbivores) and build synthetic consortia to uncover their interdependencies. We are developing new tools to construct synthetic microbial communities based on how natural anaerobic consortia interact. Our ultimate goal is to develop systems-level tools to evaluate and direct anaerobic microbial interaction, such that carbon flux can be engineered within these communities by compartmentalizing breakdown and production across microbes. Forthcoming applications of this research include engineering anaerobic digestors, anaerobic production of fuels and commodity chemicals from biomass, and identification of natural products from gut anaerobes.
Engineering Synthetic Fungal Cellulosomes as Novel Biocatalysts
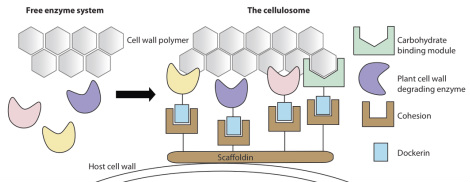
Schematic representation of a cellulosome.
Nature has evolved a powerful machine adapted for lignocellulose digestion through the use of cellulolosomes – large multi-protein complexes of tethered saccharolytic enzymes. Cellulosomes are expressed on the cell surface of anaerobic microorganisms (both bacteria and fungi) that thrive in cellulose-rich environments. In contrast to bacterial cellulosomes, fungal cellulosomes remain almost completely uncharacterized - to date the composition, architecture, and enzyme tethering mechanism remain uknown. Our group has pioneered new methods to isolate cellulosomes from diverse gut fungi to uncover the interactions and "parts" that mediate their assembly. We are now leveraging these findings to build synthetic, fungal cellulosome-inspired complexes in yeast and bacteria via synthetic biology approaches – this allows us to engineer new functionalities into the complexes, and accelerate any multi-step reaction beyond that observed in nature.
Membrane Proteins for Drug Discovery, Detection, and Diagnostics
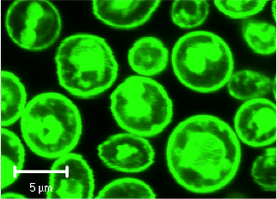
Yeast expressing GFP-tagged human adenosine A2a receptor.
The structural biology of membrane proteins has become an important focus of recent biochemical research. With at least 30% of the human genome encoding for membrane proteins, they play a vital role in cell communication and most biological functions, yet their detailed structures remain poorly characterized. G-protein coupled receptors (GPCRs) are important membrane proteins that transduce signals across cell membranes via external ligand binding, representing over 50% of all drug targets, and have been directly linked to several diseases that include HIV infection, cancer, and diabetes. While initially overlooked, oligomerization of GPCRs is now recognized to drastically impact cellular signaling, protein trafficking, and biogenesis of these membrane proteins. Our group is developing new expression and biophysical tools to isolate and study GPCR monomers and dimers to determine the factors that drive oligomerization, the stability associated with it, and how to control it in vivo. Applications of this research range from detection of low-level environmental microbial biosensors to oligomer-specific drug design.
New Membrane Proteins for Synthetic Biology
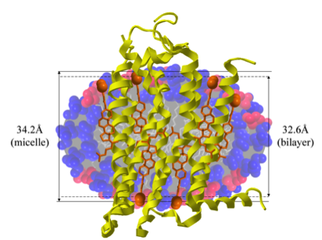
Membrane proteins are the gatekeepers of the cell, and regulate processes as diverse as selective transport to signaling. Our group is specifically focused on studying the structure-function relationship of G-protein coupled receptors (GPCRs) in eukaryotes. Though mammalian GPCRs have garnered much attention as drug targets, we have identified GPCR analogs in anaerobic fungi for the first time; these receptors play a role in sugar sensing and the reprogramming of metabolism. Our group is combining bioinformatic approaches with biochemical screening to decipher the function of these unusual gut fungal membrane proteins. By transferring this machinery to model microbes, we aim to endow them with the ability to utilize an expanded set of compounds for bioproduction and remediation.
© 2022 All Rights Reserved.